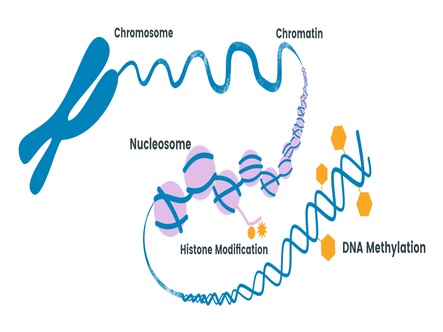
DNA Methylation in Ecology and Evolution
Dates
10th February 2025 – 14th February 2025
To foster international participation, this course will be held online
Overview
Epigenetic studies on DNA methylation are now possible to do at the nucleotide level genome-wide and also in non-model organisms. In this course, we will introduce the different available
approaches for obtaining and analysing DNA methylation data, including bisulfite sequencing (BS-seq, EM-seq) with Ilumina and long reads with Oxford Nanopore. We will also touch on PacBio
long-read analysis. We will cover all necessary steps to obtain methylation information from high throughput data to statistical analyses used to identify differentially methylated sites and
regions. The data will be interpreted in terms of their biological importance in the field of ecology and evolution.
format
The course will be delivered over the course of five days. Each day will include an introductory lecture with class discussion of key concepts. The remainder of each day will consist of practical
hands-on sessions. These sessions will involve a combination of mirroring exercises with the instructor to demonstrate a skill as well as applying these skills on your own to complete individual
exercises.
Target Audience
This course is aimed at researchers and technical workers who are or who will be generating and/or analysing DNA methylation from whole genome bisulfite sequencing data from IIlumina, PacBio data
or Nanopore data. Examples demonstrated in this course will involve primarily non-model organisms with a draft reference genome available and examples of applications of this data type for
different purposes will be covered.
program
Monday: DNA methylation analysis with short reads (Illumina WGBS and EM-Seq) – 2-7pm Berlin time
- Presentation: Origin and evolution of DNA methylation in eukaryotes: from parasite defence
to gene regulation.
- Presentation: Introduction to short-read format and processing (WGBS/EM-Seq).
- Lab 1: Short-read processing:
o QC, filtering, trimming
o Genome preparation
o Alignment, deduplication
Instructors
Cost overview

Cancellation Policy:
> 30 days before the start date = 30% cancellation fee
< 30 days before the start date= No Refund.
Physalia-courses cannot be held responsible for any travel fees, accommodation or other expenses incurred to you as a result of the cancellation.



