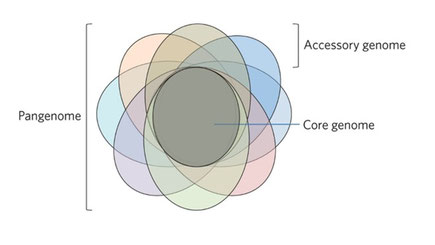
Analysis of Prokaryotic Pangenomes
Dates
ONLINE, 15-17 April 2024
To foster international participation, this course will be held online
Course overview
An increasing number of genomes are being sequenced every year, highlighting the extent of genetic variation encoded in the genomes of closely related organisms, especially prokaryotes. It is becoming clear that a single reference genome, which cannot take this variation into account is not informative enough to answer big scientific questions. Instead, genomicists are moving towards the publication and study of pangenomes – descriptions of the total genetic variation in a set of organisms. Even within single species, prokaryotes vary hugely not only in their DNA sequence but even in the genes included encoded in their genomes. In this course, you will be introduced to the study of pangenomes and their implications in biological research. You will also be given the chance to put theory into practice. By the end of the first day, you will have analysed a set of bacterial genomes to illustrate their pangenome. On day 2, you will assess the extent to which sets of genes evolve together in pangenomes and make inferences about the implications of this in a range of biological fields. Finally, in day 3, you will have the chance to design your own bespoke analysis, putting the theoretical and practical knowledge gained in the first two days into practice with the aid of the course leads.
Target audience and assumed background
This course is aimed at scientists at the level of graduate student, post-doctoral researcher or academic looking to understand the field of pangenomics. All the practicals will use the unix command line, which can be accessed via mac, linux or though the ubuntu app or a virtual machine on windows. Although not essential, it would be beneficial to have some experience of unix. In addition to this, an understanding of basic genetics, genomics and evolutionary biology will aid understanding of the concepts discussed.
Program
Monday- 10 am -6.30 pm Berlin time zone
- Lecture 1 - Welcome, Introduction to the Faculty, Introduction to the pangenome.
- Lecture 2 - Bacterial genomics from the very basic principles - what a genome is and what it looks like in data. Where to download it. Genome annotation using Prokka. BUSCO analysis of genome completeness. Command line basics
- Practical 1 - annotating genomes and checking their BUSCO completeness
- Lecture 3 - Pangenome construction – Inferring homology using BLAST. Pangenome construction tools including Roary, Pantagruel and Panaroo. Details of Panaroo output and process. Pangenome statics including gene occupancy histograms and pangenome graphs. Problems with method.
- Practical 2 – Constructing a bacterial pangenome
- Summary of the day
Tuesday- 10 am -6.30 pm Berlin time zone
- Lecture 4 - Relationships between genes in a pangenome
- Lecture 5 – Networks and their visualisation using gephi. Phylogeny and the D score.
- Practical 3 – Running Coinfinder on a bacterial pangenome.
- Practical 4 - Random Forests – predicting gene presence and absence in bacterial genomes.
- Summary of the day
Wednesday- 10 am -5.30 pm Berlin time zone
- Lecture 6 – Refresh and introduce the final practical
- Practical 5 - free practical – students design own experiment with their own data (data can be provided by organisers).
- Group discussion - Problems, strategies for analysis.


Cost overview
Should you have any further questions, please send an email to info@physalia-courses.org
Cancellation Policy:
> 30 days before the start date = 30% cancellation fee
< 30 days before the start date= No Refund.
Physalia-courses cannot be held responsible for any travel fees, accommodation or other expenses incurred to you as a result of the cancellation.
